In this tutorial we will demonstrate how to run a forward simulation with FLEXPART and how to visualise the results in various ways.
Please enter folder 'fwd' to start working.
Running the simulation
We will simulate a volcano eruption by releasing some SO2 from the Icelandic volcano Eyjafjallajökull.
The simulation itself is defined by the 'fwd_conc' FLEXPART Run icon and the 'rel_volcano' FLEXPART Release icon, respectively. Both these are encompassed in a single macro called 'fw_cond.mv'. For simplicity will use this macro to examine the settings in detail.
The macro starts with defining the release like this:
rel_volcano = flexpart_release(
name : "VOLCANO",
starting_date : 0,
starting_time : 15,
ending_date : 2,
ending_time : 12,
area : [63.63,-19.6,63.63,-19.6],
top_level : 9000,
bottom_level : 1651,
particle_count : 10000,
masses : 1000000
)
This says that the release will happen over a 45 h period between heights 1651 and 10000 m at the location of the volcano and we will release 1000 tons of material in total.
Please note that
- the species is not defined here (will be defined in
flexpart_run()) - we used dates relative to the starting date of the simulation (see also in
flexpart_run())
The actual simulation is carried out by calling flexpart_run():
#Run flexpart (asynchronous call!)
r = flexpart_run(
output_path : "result_fwd_conc",
input_path : "../data",
starting_date : 20120517,
starting_time : 12,
ending_date : 20120519,
ending_time : 12,
output_field_type: "concentration",
output_flux : "on",
output_trajectory : "on",
output_area : [40,-25,66,10],
output_grid : [0.25,0.25],
output_levels : [500,1000,2000,3000,4000,5000,7500,10000,15000],
release_species : 8,
receptors : "on",
receptor_names : ["rec1","rec2"],
receptor_latitudes : [60,56.9],
receptor_longitudes : [6.43,-3.5],
releases : rel_volcano
)
print(r)
Here we defined both the input and output path and specified the simulation period, the output grid and levels as well. We also told FLEXPART to generate gridded concentration fields and plume trajectories on output..
The actual species that will be released is defined as an integer number (for details about using the species see here). With the default species settings number 8 stands for SO2.
If we run this macro (or alternatively right-click execute the FLEXPART Run icon) the results (after a minute or so) will be available in folder 'result_fw_conc' . The computations were actually taken place in a temporary folder then metview copied the results to the output folder. If we open this older we will see two files there:
- conc_s001.grib is GRIB file containing the gridded concentration fields
- tr_r1.csv is CSV file containing the plume trajectories
Please note that these are not the original outputs form FLEXTRA but were converted to formats more suitable for use in Metview. For details about the FLEXPART outputs please click here.
Visualising gridded fields
To plot a particular parameter and level we need to filter the desired dataset from the resulting FLEXPART output file. Unfortunately, Metview's Grib Filter icon cannot handle these files (partly due to the local GRIB definition they use) so we need to use other means to cope with this task. For this reason and also to make FLEXPART output handling easier a set of Metview Macro Library Functions were developed. We will heavily use these functions in the examples below.
Inspecting the FLEXPART GRIB file
Before seeing the macro code to generate the plot we inspect the file itself we wanted to plot. Double-click on the 'conc_s001.grib' GRIB icon' in folder 'result_fw_conc' to start up the Grib Examiner. We can see that this file only contains "mdc" (=Mass density concentration) fields. We can find out further details about this parameter by setting the Dump mode to Namespace and Namespace to Parameter:
Generating the plot
The macro to visualise the concentration fields on a given level is 'plot_level.mv'.
First, we define the level (8000 m) and the parameter ("mdc") we want to plot. Then we call the Macro Library Function mvl_flexpart_read_hl() to extract the data to plot.
dIn="result_fwd_conc/" inFile=dIn & "conc_s001.grib" lev=8000 par="mdc" #Read fields on the given height level g=mvl_flexpart_read_hl(inFile,par,lev,-1,1)
Next, we define the contouring definition. The units we used here is ng m**-3 because for parameter "mdc" the native units (kg m**-3) are automatically scaled by the plotting library (see details about the this scaling for various FLEXPART GRIB fields here.
#The contour levels
cont_list=[1,10,50,100,150,200,250,500,750,1000,2000,6000]
#Define contour shading
conc_shade = mcont(
legend : "on",
contour : "off",
contour_level_selection_type : "level_list",
contour_level_list : cont_list,
contour_label : "off",
contour_shade : "on",
contour_shade_method : "area_fill",
contour_shade_max_level_colour : "red",
contour_shade_min_level_colour : "RGB(0.14,0.37,0.86)",
contour_shade_colour_direction : "clockwise",
contour_method: "linear"
)
Next, we build the title with mvl_flexpart_title(). Please note that we need to explicitly specify the plotting units!
title=mvl_flexpart_title(g,0.3,"ng m**-3")
Finally we define the view with the map:
and generate the plot:
plot(view,g,conc_shade,title)
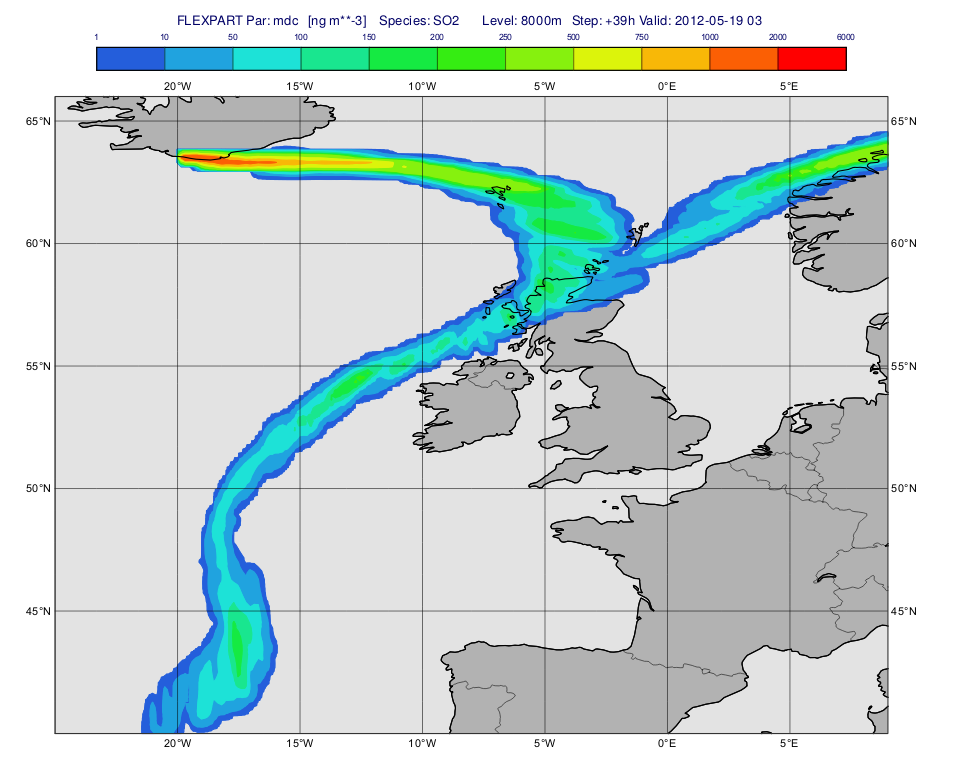
Having run the macro we get a plot like this (after navigating to step 39h):
Computing and plotting total column mass
Using the simulation results we will compute and plot the total column integrated mass. The macro to use is 'plot_total.mv'.
First, we call mvl_flexpart_total_column() to perform the computations. The result is a fieldset containing "tcmd" fields with units of "kg m**-2".
dIn="result_fwd_conc/" inFile=dIn & "conc_s001.grib" #Compute the total column integrated mass g=mvl_flexpart_total_column(inFile,1)
Next, we define the contouring. The "tcmd" fields are autocratically scaled into "ng m**-2" for contouring but with the current value range it is better to use "g m**-2" units. To achieve it we simply multiply the "tcmd" fieldset with 1000:
g=g*1000
Then specify the contour levels
cont_list=[0.0001,0.001,0.005,0.01,0.02,0.05,0.1,0.5]